
There are few diseases that match the persistence and pervasiveness of tuberculosis (TB). Mycobacterium tuberculosis, the bacterium that causes the infection, resides in the bodies of one-third of the human population. Most infections are latent, meaning that they do not produce any symptoms. In latent TB, the bacteria lie dormant in the lungs, secluded from the rest of the body and suppressed by immune cells called macrophages. Latent TB, however, can suddenly become active, causing symptoms that include cough, fatigue, and loss of appetite. In healthy patients, dormant bacteria have only a 10 percent chance of becoming active. In immunocompromised patients, however, the chance of reactivation is greatly increased. Indeed, throughout the past ten years, the number of patients infected with both HIV and TB has increased dramatically.
Tuberculosis and HIV
Geographically, TB is prevalent throughout Asia, Eastern Europe, Russia and Sub-Saharan Africa. With 24 percent of the world’s TB cases, Africa has the highest per capita mortality from the disease. The region also happens to be the epicenter of the HIV epidemic. HIV destroys the ability of the human host’s macrophages to control TB infection, a fact that explains the high lethality of the disease in patients with both TB and HIV compared to TB in otherwise healthy individuals. This fact has exacerbated the global TB epidemic, especially in regions of the world where TB and HIV are most prevalent.
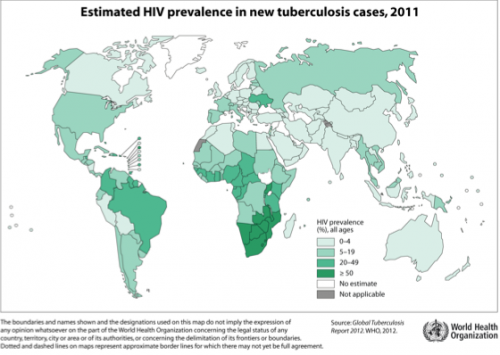
Why Sub-Saharan Africa?
While the despairingly high prevalence of TB and HIV in Sub-Saharan Africa could once have been attributed to the lack of public health initiatives and infrastructure in the region, researchers have now pinpointed a more basic reason for the disease’s ethnocentricity (meaning its high incidence among specific populations). Our genetic makeup has long been known to play a role in our susceptibility to tuberculosis and its progression. Researchers including Dr. Richard Bucala, Professor at the Yale School of Medicine, are examining the genomes of different human populations in an attempt to identify small differences that might make certain populations more susceptible to diseases such as tuberculosis.
Specifically, Bucala’s research investigates how genes can influence the body’s response to different pathogens. Bucala’s lab recently discovered a functional polymorphism in a gene that strongly influences our response to TB. This gene is called Macrophage migration inhibitory factor (MIF), and the immune protein it encodes is produced by agents of the immune system (macrophages, leukocytes, and pulmonary epithelial cells) upon infection. The MIF gene has a common polymorphism in its promoter region that results in two main variants: a low-expressing variant and a high-expressing variant. The difference between these variants, or alleles, lies in the different number of short DNA repeats in the promoter. The low-expressing variant has five repeats of a tetranucleotide regulatory sequence while the higher-expressing variants have more than five repeats, leading to correspondingly greater expression. After discovering this polymorphism, Bucala started measuring the distribution of these variants in human populations within the United States.
In nursing home studies carried out previously by other researchers, African-American patients had been shown to be more susceptible and to develop more lethal tuberculosis than Caucasian patients. Interestingly, Bucala found that while the low-expressing variant is expressed by 65 percent of African Americans, it is only expressed by 45 percent of Caucasians. Noticing this trend, Bucala then sequenced the MIF gene in different populations from around the world. The low-expressing variant was most prevalent in Sub-Saharan Africa, with 78 percent of Zambians, for instance, possessing the low-expressing variant.
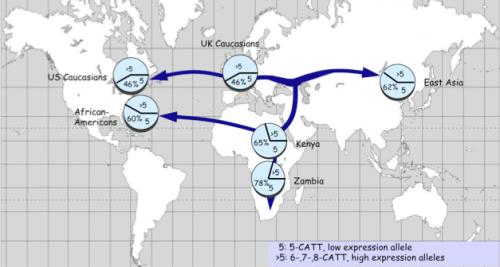
MIF and Malaria
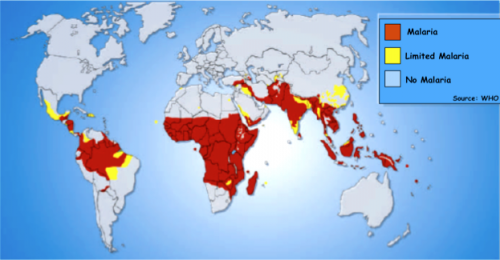
The prominence of the low-expressing MIF variant was not the only genetic discovery within the Zambian population. Unlike their counterparts in Northern African and Mediterranean countries, populations in Southern Africa were known to lack the sickle cell gene, a variant in the hemoglobin gene that conferred malarial immunity in heterozygous individuals at the cost of anemia and shortened lifespan in homozygous individuals. Realizing this, Bucala hypothesized that the low-expressing MIF gene helps provide immunity against malaria, as it suppressed the excessive inflammatory response that often kills malaria-infected children. The group studied African children with malaria and found that a low-expression MIF allele was associated with less severe disease.
Mouse models of malaria or TB infection performed by Bucala confirmed that having a low-expression variant of the MIF gene correlates with less severe malaria but a higher likelihood of developing TB pathology. Accordingly, it was predicted that TB patients with low MIF would be more susceptible and less able to fight off TB infection. Rita Das, a graduate student and infectious disease fellow in the Bucala lab, found evidence for this in humans as HIV-positive Ugandan patients with disseminated (widespread) TB were 2.4 times more likely to have a low-expressing variant of the MIF gene than those without bacteremia.
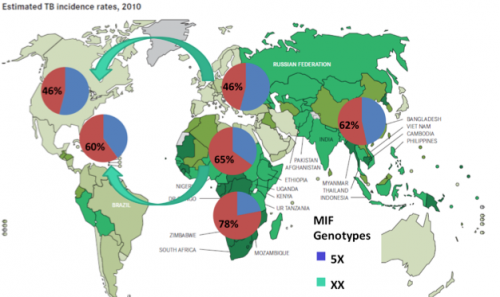
The low-expressing form of the MIF gene is clearly a double-edged sword. The low-expression variant, by preventing a full-blown immune response, protects individuals from the lethal inflammatory complications of malaria. At the same time, however, the variant can increase susceptibility to TB and act as a marker for poor prognosis of disease progression.
Population Stratification of the MIF locus
Low-expressing MIF alleles occur disproportionally in Africa. Bucala postulates that this stratification of the MIF locus occurred when a small population of individuals migrated out of Africa about 100,000 years ago. As this population reached colder climates with less malaria and encountered new, life-threatening pathogens, the benefit of the low-expressing allele was lost in favor of the higher-expressing allele.
You Take the High Road and I’ll Take the Low Road
While the prevalence of the low-expressing form of the MIF gene in Sub-Saharan Africa can be explained by the immunity it confers to lethal malaria, its prevalence in areas such as North America and Europe cannot be explained as easily. Common diseases in these regions include community-acquired pneumonia, a disease that can lead to sepsis (a state of overwhelming infection) and death if not treated. Community-acquired pneumonia is the leading cause of non-cardiac intensive care unit deaths in the U.S., simply because of how easily sepsis can be triggered. The low expressing MIF allele has been shown to not only exacerbate TB in African populations, but to lead to reduced survival rates from sepsis in Americans with community-acquired pneumonia. In these cases, the high-expressing allele is clearly beneficial.
Bucala’s clinical expertise is rheumatology, particularly dealing with the role of the MIF gene in autoimmune diseases such as arthritis. These diseases are caused by excessive inflammation, highlighting an important context for favoritism of the low-expressing allele in North America. Studies have shown that the low-expressing form of MIF is associated with less severe rheumatoid arthritis, systemic lupus erythematous, and other inflammatory disorders. Genes that have arisen to combat different infections also have the ability to worsen autoimmune inflammation, emphasizing the balancing act that our immune system plays to keep us healthy.

Individualized Treatment
Knowing what variant of the MIF gene a person has can lead to a greater understanding of that person’s disease, and how best to treat it. Those suffering from TB and having a low-expression MIF allele theoretically could receive supplemental MIF to reset their immune response toward one that can combat TB more effectively. However, as is the case with many infectious diseases, the epicenter of the HIV/TB dual epidemic is in a part of the world where analysis of each person’s genome may not be cost-effective. First, rapid and inexpensive techniques are needed to determine each individual’s MIF variant.
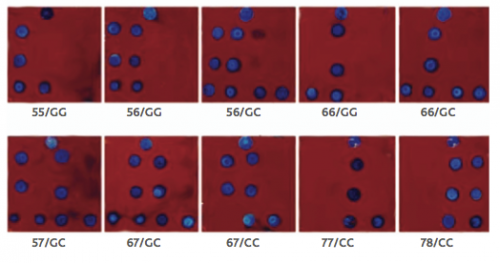
To this end, rapid diagnostic methods have been developed using biochips. These are small silicon chips with oligonucleotides attached to the surface, each of which can light up when bound to a specific sequence. Using this technology to detect which patients are more susceptible to malaria or TB infection will allow doctors to spend more time with their cases, whether that involves longer hospitalizations, more antibiotic use, or an individually optimized treatment plan.
According to Bucala, “we are slowly developing new antibiotics for TB, but we desperately need new tools to combat this global epidemic.” Bucala is currently working with Dr. William Jorgensen in the Yale Department of Chemistry on ways to supplement the MIF gene product in low MIF-expressing individuals. Supplementing the gene product would provide an important adjunct to the current method of combating resistant TB strains, which involve taking multiple antibiotics for many months, a difficult and costly treatment and one being circumvented by the ever-increasing drug-resistance of TB strains.
The discovery of geographic and ethnocentric polymorphisms in the MIF gene has led to a new understanding of the HIV/TB dual epidemic. No longer is geography simply an additional fact about the epidemic; it now offers a unique explanation of its cause.
About the Author: Emma Graham is a sophomore in Trumbull College majoring in Molecular Biochemistry and Biophysics.
Acknowledgements: The author would like to thank Dr. Richard Bucala for taking the time to share his research.
