If you’re hoping to run a fruitfly drinking contest, you should definitely invite the species Drosophila melanogaster (though you might need some really tiny cups). This species of fruitfly, which lives in alcohol-rich rotting fruits in the wild, is able to tolerate a far higher environmental alcohol concentration than its close relatives. Previously-done gene sequencing analysis suggested that one reason for the difference is that Drosophila melanogaster has a slightly different alcohol dehydrogenase (ADH) protein, an enzyme responsible for breaking down alcohol, than its close relatives.
Within the evolutionary biology community, there is a longstanding hypothesis explaining the high alcohol tolerance of Drosophila melanogaster: in order to adapt to its ethanol-rich environment, this species’ ADH protein has evolved to become more efficient at breaking down alcohol inside cells. The explanation sounds fairly intuitive, but researchers recently found evidence against this hypothesis. The research team, working at the University of Chicago, genomically engineered Drosophila to produce the ADH gene of its two million-year-old ancestors—which were almost certainly less tolerant of alcohol—and discovered that the ADH gene did not evolve to become more efficient at breaking down alcohol.
“This research addresses a major challenge in evolutionary biology,” said Mo Siddiq, lead scientist on the project. “Not only can we build hypotheses based on correlated observations, we can actually go back in and test them biologically. That has historically been the hardest part.”

Siddiq’s research has its roots in the history of how the field of evolutionary biology was born; theories about evolution have been in existence since ancient times. To the Ancient Greeks, Romans, and Chinese, the study of the relationships between and development of species was considered a branch of philosophy. Modern evolutionary biology began to develop in the late 19th and early 20th centuries, when Charles Darwin’s theory of natural selection and Gregor Mendel’s laws of biological inheritance were resolved in a joint mathematical framework.
In the mid-20th century, there was a huge breakthrough in molecular biology when DNA was discovered to be nature’s hereditary material and researchers developed the ability to detect differences in protein and ultimately DNA sequences. These new molecular approaches provided important tools for studying evolution, but they led to a splintering in many biology departments, with molecular biologists and biochemists falling on one side and evolutionary biologists and ecologists falling on another. Dubbed the “molecular wars,” this conflict’s effects are still present today.
Collaboration between evolutionary biologists and biochemists is not very common, perhaps because of these historical differences. However, many scientists in both fields are working to bridge the gap. “People are starting to realize that by taking a multi-disciplinary approach, we can get an overall deeper understanding,” Siddiq said.
A key, overlapping topic for these two fields is the search for genetic changes in a species that adapt it to its particular environment. Genes dictate how the organism functions, just like a recipe tells chefs how to make food. If the customer isn’t happy with a dish, the chef might swap the recipe, forever changing how that dish is made in the future. The same is true of natural selection on genes, except nature selects for genetic changes that make organisms better capable of surviving and reproducing in their environments (and natural selection takes a lot longer) . Oftentimes, distinct patterns of DNA sequence, called signatures of selection, are produced when natural selection is acting or has acted on a gene, and this can point the way towards genetic changes that might have been important in the evolutionary history of a species. However, many other factors, such as random mutations, can also lead to rapid gene changes during evolution, and some scientists remain cautious about interpreting what signatures of selection mean for biological functions.
Siddiq and his advisor Joe Thornton set out to test the direct link between genetic changes in ADH and whether those changes actually played a role in adaptation of D. melanogaster to alcohol-rich environments. They chose this system because there was a clear hypothesis of cause and effect, and the genetic tools available in the fruitfly, a widely-studied model in evolutionary biology, made it easily-testable.
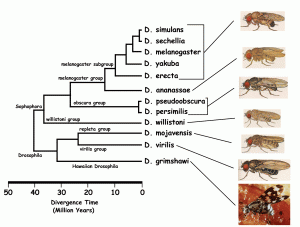
Drosophila melanogaster evolved to survive in ethanol-rich environments after it split from its sister fly species, Drosophila simulans, between two to four million years ago. So, the scientists set out to determine the ADH gene sequence from the last common ancestor between Drosophila melanogaster and Drosophila simulans. Using statistical methods, they traced back the gene sequence of ancient ADH by comparing the genetic sequence of modern Drosophila melanogaster to that of its modern relatives.
With the ancient ADH sequence in hand, Siddiq then biochemically synthesized, expressed, purified, and characterized the ancient ADH protein. To do this, the gene sequence for ancient ADH was cloned into a plasmid, a small circular piece of DNA that is easily inserted into E. coli bacterial cells. The bacteria then multiplied, replicating their plasmids with each cell division and producing large amounts of the protein. The expressed ADH protein was isolated and tested for its ability to process ethanol. The result was stunning: modern ADH and ancient ADH have the same ethanol-processing ability.
To further verify this result and to test how the ancient protein functioned in the context of the actual organism, Siddiq partnered with David Loehlin from the University of Wisconsin and Kristi Montooth from the University of Nebraska to recreate living Drosophila melanogaster fruit flies that express the ancient ADH gene. The team first produced DNA coding for the ancient ADH sequence, then injected this DNA into fruitfly eggs. They bred thousands of these “ancestralized” flies, and tested their ability to process ethanol by putting them in chambers filled with different ethanol concentrations. This result mirrored the previous one: the ancestralized flies were just as good at metabolizing ethanol as their modern counterparts. The scientists concluded that the species’ enhanced ethanol-processing ability is not a result of adaptation in the ADH protein, despite the signature of selection on ADH gene sequence during evolution. This suggests that the previous hypothesis—which assumed that changes in the Drosophila gene sequence were an environmental adaptation—may be wrong, and other genes need to be studied to understand how alcohol tolerance evolved.
But why might a generation of evolutionary biologists have been wrong all this time about the role of ADH? The main reason could be that alcohol tolerance is complex, and dozens of different genes not directly involved in alcohol metabolism might have played an important role.
Unlike college students, most biologists don’t care much about how “buzzed” fruit flies can get. However, fruit flies are a central model species that molecular biologists and geneticists use to test their theories. This research could overturn a long-standing hypothesis in evolutionary biology and show us a new way with which hypotheses about the past can be tested. Moreover, the study verified the utility of newly-developed tools in molecular biology that have a wide-ranging impact for future research.
Ultimately, this research only came through an incredible collaboration between a biochemist in Chicago, a geneticist in Wisconsin and a physiologist in Nebraska. These three individuals had met throughout their academic careers, at conferences and next-door labs, and developed a connection that led them to an interdisciplinary approach for testing their scientific hypotheses—an approach that would have never come from a single department. Perhaps this bodes well for young biologists: like the humble fruit fly, they can go out with their colleagues and get some drinks, all the while developing novel hypotheses in evolutionary biology.

