Image courtesy of Wikimedia Commons
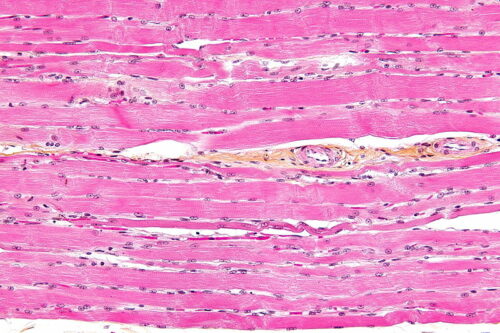
Muscular Dystrophy
Duchenne muscular dystrophy (DMD) is a genetically linked disorder caused by a mutation of the protein dystrophin. The dystrophin protein transfers muscle contractions from inside of a muscle cell outward. In other words, it connects the center of a muscle cell outwards. Without functional dystrophin proteins, muscle cells are not intact, leading to muscle degeneration and weakness, and eventually bringing on the loss of heart and skeletal muscles. With no cures for DMD, an optimistic life expectancy for those with DMD is in the early thirties.
Anton Bennett, Dorys McConnell Duberg Professor of Pharmacology and Professor of Comparative Medicine at Yale, and his team discovered a possible treatment pathway for DMD. They found this pathway while researching mitogen-activated protein kinases and its phosphatases.
How the Cell Propagates Information
Mitogen-activated protein kinase (MAPK) are a family of proteins that play an important role in signal cascades, a series of reactions that occur one after the other, responding to an activation event. Like all proteins, MAPKs consist of a chain of amino acids, or residues, which fold into specific conformations based on the interactions between the different types of amino acids. MAPKs are activated by the phosphorylation of tyrosine and threonine residues, which are located on the activation loop of the protein. Phosphorylation involves adding a phosphate group to the protein, often activating a function. This activation is often in response to an outside signal, such as a ligand binding to a signal receptor within the cell membrane of a cell. After activation, MAPKs phosphorylate subsequent substrates (the targets of an enzyme), initiating a signal cascade. These signal pathways elicit a large range of cellular responses such as gene transcription, cellular differentiation, metabolism rate, apoptosis and more.
Mitogen-activated protein kinase phosphatases (MKPs) offer an additional layer of control by regulating MAPKs. They do this by dephosphorylating the tyrosine and threonine residues at the activation loops. Without phosphate groups, MAPK return back to their inactive forms. MKPs are thought of as belonging to different sub-families, each of which regulates a specific group of MAPKs. For example, a group of MKPs which includes MKP5, MKP7, and DUSP8 targets stress-activated MAPKs. This specificity allows MKPs to closely regulate their respective kinase proteins, providing the cell finer control of its molecular machinery.
The Undruggable MKP5
While MKPs are essential for signal regulation, recent research suggests that they may also have roles in disorders. Specifically, studies suggest that MKP5 disrupts muscle myogenesis, or the formation of muscle tissue. When skeletal muscle cannot regenerate through myogenesis, they are replaced with fibrotic, or connective, tissues.
In this sense, MKP5 seems to be the perfect target to treat this disorder. However, the phosphatase is tricky to inhibit as most identified inhibitors attack the active site of the phosphatase, which is positively charged. Like the attraction between the north and south poles of a magnet, to attach to the positively charged site, the inhibitor compounds are often negatively charged. However, charged molecules are undesirable for drug use as these molecules cannot pass through the cell membrane. In light of these challenges, many deemed MKP5 to be “undruggable.” Furthermore, inhibiting a member of the MKP family is further complicated by the similarity of phosphatase structures. A molecule that inhibits MKP5 may also inhibit other MKP molecules, disrupting other mechanisms within the body. Thus, the specificity of the molecule adds an additional layer of difficulty when developing a suitable inhibitor.
To target MKP5, Bennett implemented a novel strategy: to look for inhibitors that could deactivate the enzyme without targeting the active site, also known as allosteric inhibitors. “That was really the breakthrough strategy,” Bennett said.
Screening
The first step to finding inhibitor molecules was to test a large set of molecules to determine if any of them loosely inhibited MKP5. There are two main ways to screen for molecules: physically or computationally. “We did a real, physical screen where we screened over a hundred thousand compounds using automation with [many] compounds in well plates,” said Zachary Gannam, a postdoctoral fellow and the first author of the study published in Science Signaling. Alternatively, virtual screenings use structural biological information to create docking sites on a desired protein. Then, simulations can dock millions of compounds to test their interactions.
Both screening methods have their benefits and drawbacks. Physical screening requires a significant amount of protein, an optimized experiment, and is also more expensive and time intensive than computational methods. However, when you get a hit, you know the compound is an inhibitor. On the other hand, computational screenings require researchers to have an idea of the protein structure and the structure of the sites where small molecules can interact. After conducting computational models, researchers are required to eventually physically test the compounds with the most promise. A positive is not necessarily guaranteed to be an inhibitor, and the highest hit rates are still fairly low. With insufficient information about docking sites on MKP5 for computational screenings, the research team opted to conduct physical screenings. The screening tests revealed that a molecule, denoted by Compound 1, showed promise in inhibiting MKP5.
The How’s and the Why’s
With initial screenings showing the inhibitory properties of Compound 1, Bennett’s team wanted to find out how the molecule interacts with MKP5. Understanding the molecular interactions between MKP5 and Compound 1 required acquiring a crystal structure—a repeated lattice of stable protein interactions—of the two interacting molecules, which would help researchers determine the structure of the proteins and their interaction. “This was the first crystal structure of an MKP in a complex with a small molecule,” Gannam said. The lack of previous procedures for crystallography meant that the team had to test out the MKP5-Compound 1 complex in solutions of various combinations of buffers, salts and precipitants. “It’s very idiosyncratic and there are not many set rules to follow,” Gannam said. Rounds and rounds of screening for crystallization were required to finally develop the crystal structure.
The structure revealed that Compound 1 fundamentally shifts the shape of MKP5. Notably, a distinct allosteric site on the protein shifts to interact with Compound 1. These shifts cause the volume of the active site to decrease by eighteen percent. Analyzing the specific residues which Compound 1 interacts with also showed its selectivity for MKP5 as opposed to other MKPs within the molecule family. Specifically, the research team showed that methionine and threonine residues on MKP5’s allosteric site were unique to it. Further tests revealed that Compound 1 was less effective at inhibiting a mutated MKP5 with altered methionine and threonine residues. Thus, Compound 1 seems to selectively bind to MKP5 due to these two residues.
Moving to Cells
So far, research on Compound 1 had been conducted in vitro, or outside of the cell. To make sure that Compound 1 behaves predictably within a biological context, the team investigated the effect of Compound 1 in mice cells. Since MKP5 inhibits MAPK and JNK, introducing Compound 1 would inhibit MKP5, therefore increasing the phosphorylation of MAPK and JNK. Not only did Compound 1 increase MAPK and JNK activities, it had no effect on other kinases such as ERK1/2, which is not regulated by MKP5. These results showed that even in a cellular context, Compound 1 seems to only inhibit MKP5, displaying the specificity that is crucial for viability as a drug.
What’s Next?
Discovering Compound 1 represents the crucial first step to developing a treatment for DMD. However, there is still a long way to go to produce a viable drug. “Essentially, we have a drug development project to make a compound that is ideally highly potent, orally viable and fits the once-a-day pill treatment,” Bennett said.
Compound 1 may also have applications beyond DMD. Compound 1 targets the pathway leading to tissue fibrosis or the thickening of scarring on tissue. For example, postoperative fibrosis is a type of complication that occurs after surgeries, involving excess tissue scarring as a result of the surgery. “Fibrosis accounts for forty-five percent of deaths worldwide in various clinical presentations,” said Bennett. These include cardiac, lung, liver, and kidney fibrosis. Compound 1 can potentially address these fibrosis diseases and complications.
The path to a workable drug requires meticulous and thorough work. It allows for innovations like Compound 1 to have the potential to help millions.
References
AM;, M. (n.d.). Loss of MKP-5 promotes myofiber survival by activating STAT3/Bcl-2 signaling during regenerative myogenesis. Retrieved November 26, 2020, from https://pubmed.ncbi.nlm.nih.gov/29047406/
Bennett, A. M. (2019, January 01). MKP5 in Dystrophic Muscle Disease. Retrieved November 26, 2020, from https://grantome.com/grant/NIH/R01-AR066003-05
Gannam, Z. T., Min, K., Shillingford, S. R., Zhang, L., Herrington, J., Abriola, L., . . . Bennett, A. M. (2020). An allosteric site on MKP5 reveals a strategy for small-molecule inhibition. Science Signaling, 13(646). doi:10.1126/scisignal.aba3043
Zhang, W., & Liu, H. (n.d.). MAPK signal pathways in the regulation of cell proliferation in mammalian cells. Retrieved November 26, 2020, from https://www.nature.com/articles/7290105