Imagine a world without DNA.
That is, a world before DNA. Despite our fascination with the double helix and its role as the crucial core of all living things, many types of living organisms existed before DNA did. Indeed, it is probable that life began with what is now considered DNA’s little sidekick, the supporting actor of the molecular world: ribonucleic acid, or RNA.

The RNA World
The idea of RNA as the ancestor of modern life is not new. It arose in the mid-20th century, in the papers of scientists like Francis Crick and Leslie Orgel. Termed the “RNA World Hypothesis,” it proposes that almost four billion years ago, the world was teeming with organisms controlled by ribonucleic acids — that is to say, organisms with no DNA at all. In this world, single-stranded RNA molecules stored the genetic information and acted as catalysts for chemical reactions. Since RNA-powered organisms were self-replicating and had the ability to evolve, they met the Darwinian standards for living things. The RNA World these organisms populated would eventually evolve to include the molecules we know today, like proteins and DNA.
At its opening eighteen years ago, Yale Professor Ronald Breaker’s lab began investigating the complex functions of nucleic acids, the building blocks of RNA. Then the RNA World Hypothesis spurred the lab in a new direction.
“The RNA World Theory, regardless of how strong one thinks it is,” says Breaker, “certainly inspired me and inspired the lab to try to push nucleic acids into areas that we thought biology perhaps either had never done before, or had done four billion years ago.” Guided by the hypothesis that modern life developed from RNA organisms, Breaker began investigating ribozymes. These are RNA molecules that would have substituted for modern-day proteins in an RNA World: they are catalysts, speeding up chemical reactions and enabling them to proceed. The lab managed to create synthetic ribozymes in test tubes and discovered more about how they work.
In manufacturing ribozymes, Breaker’s lab began exploring the limits of RNA. Could manipulation of nucleotide building blocks yield previously unknown structures and functions for RNA? In creating new molecules in test tubes, he had a breakthrough: he was able to build an RNA molecule with the ability to flip genes on and off like a light switch. RNA molecules with this function are called riboswitches, and their discovery led to a new and very important understanding of RNA’s capabilities and history.

Genetic Light Switches
For decades, molecular biologists considered huge portions of DNA to be “junk” regions, because they do not contain genes that code for specific proteins. Because a strand of RNA starts out as an exact mirror copy of one of a double helix’s strands, RNA too contains these regions of so-called meaningless nucleotides. Before a newly minted RNA molecule leaves the nucleus, the “useless” parts are cut out and the useful coding regions are spliced together. The shortened RNA molecule is then free to proceed into the cell, where its instructions are parsed by a ribosome and a protein is built.
Riboswitches fit into this process of gene expression in the phase before the RNA leaves the nucleus. They are structures located in the noncoding regions close to certain genes, called the 5’ untranslated regions, and their job is to bind to small molecules called metabolites (for example, glutamine or thiamine pyrophosphate). Each riboswitch selectively binds a different metabolite, and then changes its own three-dimensional folded structure upon binding. This change in molecular structure can turn a gene on or off.

In eukaryotes, the specific mechanism by which the riboswitch can shut down a gene involves spliceosomes, the bundles of small nuclear RNAs and protein factors (snRNPs) that cut out the noncoding regions of an RNA strand before it leaves the nucleus. When a riboswitch binds thiamine pyrophosphate, for example, the molecule refolds to form a new three-dimensional structure. It becomes stabilized in that shape, as if held in position by the metabolite. The sites along the RNA strand at which the riboswitch has been stabilized by a metabolite attract spliceosomes, while the sites at which the riboswitch has not been stabilized do not. By regulating where the splicing of the messenger RNA occurs, riboswitches control which instructions from which genes are put into action.
There are several known mechanisms by which riboswitches control gene expression. Besides controlling where a messenger RNA strand is spliced, they can also control whether a ribosome in the cell is able to interpret the RNA’s instructions by selectively masking or displaying the information the ribosome needs in order to bind to the RNA. In addition, riboswitches may prevent a gene from being transcribed from DNA to RNA in the first place.
The History of Life
Breaker’s findings about riboswitches and confidence in the RNA World Hypothesis have given him a clearer picture of the evolution of life. Assuming that the world began with a strand of nucleotides that formed an RNA molecule, as Breaker explains it, things quickly became competitive. The first RNA molecule to find a way to self-replicate was a happy organism, limitlessly reproducing with all the nucleotides floating around the Earth as food, until it mutated. Then competition between varying forms of self-replicating RNA began. At some point, the nucleotide building blocks, like food in any environment, became scarce, and the RNA molecules that evolved a method to disassemble neighbors into their component nucleotides were the ones that reproduced. As a result of this mutation, other RNAs began evolving defenses against the disassembling RNAs, and the competition became stiffer and stiffer until the world was brimming with complex organisms. These organisms required the ability to sense their surroundings and respond to them by modifying their own gene expression, and this is where riboswitches come into play: they were crucial devices for facilitating the development of complicated organisms. At some point, the ribosome, a complex structure made out of RNA pieces, evolved, and with it, the ability to construct proteins from amino acids.
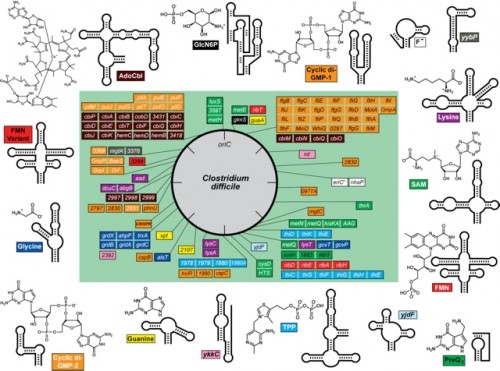
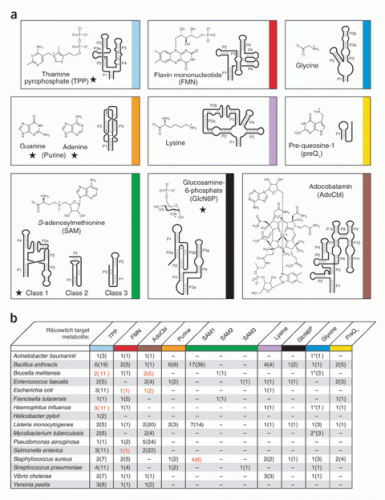
So far, the Breaker Lab has isolated and named 30 classes of riboswitches, each distinguished by a distinct tertiary structure and its own binding metabolite. All these classes have been discovered in bacteria. Every time a new class is isolated, Breaker checks the other domains of life to see whether the riboswitch also occurs there. A few have been found in eukaryotes, but so far only in plants and fungi, not humans. Breaker suspects that riboswitches exist in humans but are simply more difficult to isolate.

The lack of evidence for riboswitches in humans thus far does not mean that this research lacks implications for us. On the contrary, Breaker’s research could aid the medical field immensely. As antibiotics lose their effectiveness, we are faced with the problem of treating bacterial infections that show more and more resistance to drugs. By isolating a riboswitch that turns off a gene in a bacterium and determining the metabolite which triggers the switch, Breaker has been able to create fake metabolites to take the place of the natural ones. In other words, he has been able to create a substance that, when introduced to a bacterial cell, finds a riboswitch and turns off a gene. Because of the specificity of riboswitches to the genes they regulate and the specificity of the metabolite that triggers each riboswitch, it is possible to finely control the target of such a drug. Importantly, Breaker has found and tested a way to treat illnesses caused by bacteria such as Clostrodium difficile, which are widespread in hospitals and kill about 30,000 Americans every year. Perhaps riboswitches point to the direction medicine must follow in order to keep up with the fast pace of the changing molecular world.
While the functions of RNA are still important today, the modern world exhibits relatively little dependence on them. This is because after proteins were first synthesized, they went on to conquer the molecular world with their incredible array of possible functions. But they left in their wake riboswitches hidden within the genomes of some organisms, “relics,” as Breaker calls them, from a long-gone RNA World. These evolutionarily durable switches surely have more to tell us about the history of life, and perhaps also its future.
About the Author: Julia Rothchild is a freshman in Timothy Dwight College.
Acknowledgments: The author would like to thank Professor Ronald Breaker for his time, helpfulness, and enthusiasm for his research.